Figure 14 (page 31):
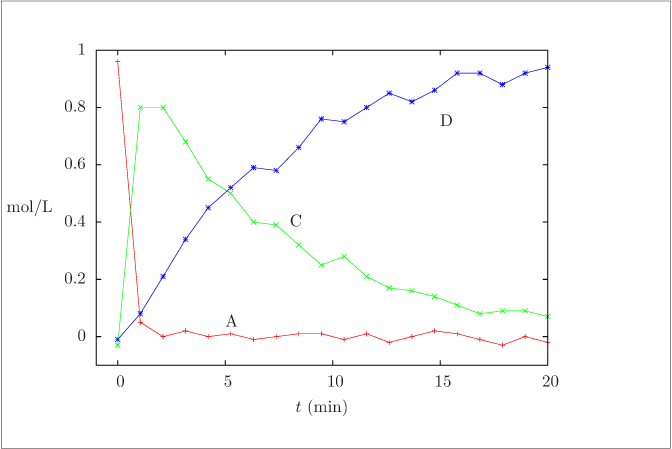
Measurements of species concentrations versus time for Exercise\ref {exer:ABCD}.
Code for Figure 14
Text of the GNU GPL.
main.m
clear('all'); close('all');
%
% Create data for the model discrimination exercise
%
% True mechanism:
%
% k1
% A+ B <-> C
% k_1
%
% k2
% C + B -> D + B
%
% jbr, 11/2008
clear all
close all
global theta
k1 = 0.25;
k_1 = 0;
k2 = 0.016;
ca0 = 1;
%cb0 = 3;
cb0 = 10;
cc0 = 0;
cd0 = 0;
c0 = [ca0; cb0; cc0; cd0];
theta = [k1; k_1; k2];
tfinal = 20;
nplot = 100;
measure.states = [1, 3, 4];
measure.time = linspace(0, tfinal, 20)';
%create the measurements by solving the model and adding noise
[tsolver, conc] = ode15s(@massbal, measure.time, c0);
randn('seed',0);
y = conc(:,measure.states);
measure.data = y + 0.02*randn(size(y));
myfile = fopen('ABCD_data.dat', 'w');
for i = 1: size(y,1)
fprintf(myfile, '%8.2f', measure.time(i), measure.data(i,:));
fprintf(myfile, '\n');
end
fclose(myfile);
figure(1);
plot(measure.time, measure.data, '-o')
massbal.m
function xdot = massbal(t, x)
global theta
ca = x(1);
cb = x(2);
cc = x(3);
cd = x(4);
k1 = theta(1);
k_1 = theta(2);
k2 = theta(3);
r1 = k1*ca*cb - k_1*cc;
r2 = k2*cb*cc;
xdot = [-r1; -r1; r1-r2; r2];
